A walk-thru our recent PennAI paper

This post is a quick walk-thru of our recent paper, “Evaluating recommender systems for AI-driven biomedical informatics.”1
Motivation
While artificial intelligence (AI) has become widespread in health care, many commercial AI systems are not yet accessible to individual researchers nor the general public due to the coding and ML expertise required to use them. We believe that AI has matured to the point where it should be an accessible technology for everyone. The ultimate goal of this project is to deliver an open source, user-friendly AI system that is specialized for machine learning analysis of complex data in the biomedical and health care domains.
Main Results
Towards that end, we developed PennAI as a browser-based tool that allows users to upload datasets and hand over analysis to an automated agent (the AI). During this process, we found that recommender systems, commonly used to recommend products to users, could meet our goals quite well for this platform. In short, PennAI is able to do the following:
- It allows users to run their own experiments or hand over analysis to an AI. The AI learns in real-time based on the user’s, and its own, experiments.
- It allows users with no programming knowledge to generate reproducible models and Python code with a few clicks.
- It generates classification results that are on-par with state-of-the-art ML systems, including in real-world application to septic shock prediction in critical care patients.
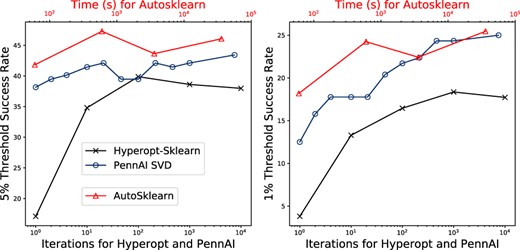 Using a simple SVD recommender, PennAI succesfully finds ML pipelines within 1\% of optimum at a similar or better rate than other AutoML tools.
Using a simple SVD recommender, PennAI succesfully finds ML pipelines within 1\% of optimum at a similar or better rate than other AutoML tools.
Methods
We were interested in evaluating recommender systems for a couple reasons. For one, they allow the user and AI to interact in lots of ways - the AI can learn from the user, and the user can be served recommendations for algorithm configurations for their dataset. In addition, recent results in AutoML23 have shown that recommender systems based on matrix factorization perform pretty well.
To evaluate different recommender systems, we took the Surprise Library and edited it to work as an online recommender system (see my fork). We then benchmarked several different recommendation strategies, including neighborhood strategies, matrix factorization, and metalearning approaches. Here metalearning is meant to refer to algorithms that use additional information about the datasets to make recommendations - in other words, we calculate a bunch of metafeatures of the datasets and compare these to decide which algorithm to run for a particular dataset.
A lot of this experiment got pushed to the supplemental material, but basically, over several experimental treatments we found that singular value decomposition (SVD) worked well for recommending an algorithm for a datset that performed as well as the best known algorithm performance for that dataset.
We were able to make those kinds of insights by leveraging a large dataset repository (PMLB)4 and by leveraging previous work we did to benchmark ML methods from Scikit-Learn5.
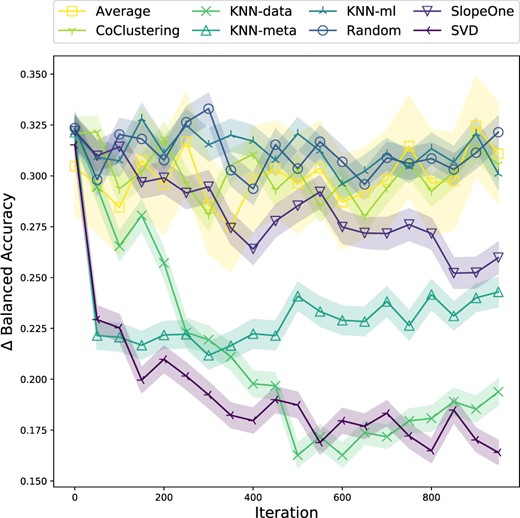 Among recommender systems, the SVD algorithm tends to converge quickest.
Among recommender systems, the SVD algorithm tends to converge quickest.
Looking Ahead
PennAI is available totally open-source and free, so we hope it will be useful to many researchers. We also hope that it will attract AutoML researchers who wish to plug in their own algorithms and use PennAI as a platform for testing their ideas. We have a sklearn wrapper in the works that will make this a little easier. It is also worth noting that we generated these results without including any feature preprocessing or engineering steps outside of core ML algorithms (and still performed well!). So there is a lot to do in terms of expanding the ML pipelines that PennAI can optimize.
If you’re interested in contributing, we’d be happy to have you!
Acknowledgments
This work was supported by members of the Computational Genetics Lab at UPenn. The development of this tool was supported by National Institutes of Health research grants (K99 LM012926-02, R01 LM010098 and R01 AI116794) and National Institutes of Health infrastructure and support grants (UC4 DK112217, P30 ES013508, and UL1 TR001878).
References
-
La Cava, W., Williams, H., Fu, W., & Moore, J. H. (2020). Evaluating recommender systems for AI-driven biomedical informatics. Bioinformatics. Oxford (open access), arXiv ↩
-
Fusi, N., Sheth, R., & Elibol, M. (2018). Probabilistic matrix factorization for automated machine learning. Advances in Neural Information Processing Systems, 3348–3357. ↩
-
Yang, C., Akimoto, Y., Kim, D. W., & Udell, M. (2018). OBOE: Collaborative Filtering for AutoML Initialization. arXiv:1808.03233 ↩
-
Olson, R. S., La Cava, W., Orzechowski, P., Urbanowicz, R. J., & Moore, J. H. (2017). PMLB: A Large Benchmark Suite for Machine Learning Evaluation and Comparison. BioData Mining. arXiv ↩
-
Olson^, R. S., La Cava^, William, Mustahsan, Z., Varik, A., & Moore, J. H. (2017). Data-driven Advice for Applying Machine Learning to Bioinformatics Problems. Pacific Symposium on Biocomputing (PSB). ^: contributed equally. arXiv ↩